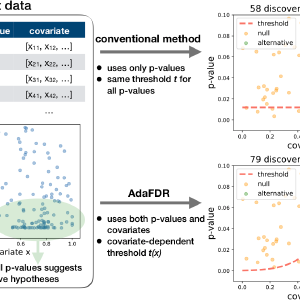
In multiple hypothesis testing, the data for each test boils down to a p-value while additional information is often available for each test, e.g., the functional annotation for genetic testing. Such information may inform how likely the small p-values are to be due to noise. Covariate-adaptive multiple testing methods utilize such information to improve detection power while controlling false positives.
Multiple hypothesis testing is an essential component of modern data science. Its goal is to maximize the number of discoveries while controlling the fraction of false discoveries. In many settings, in addition to the p-value, additional information/covariates for each hypothesis are available. For example, in eQTL studies, each hypothesis tests the correlation between a variant and the expression of a gene. We also have additional covariates such as the location, conservation, and chromatin status of the variant, which could inform how likely the association is to be due to noise. However, popular multiple hypothesis testing approaches, such as Benjamini-Hochberg procedure (BH) and independent hypothesis weighting (IHW), either ignore these covariates or assume the covariate to be univariate. We develop covariate-adaptive multiple testing methods, NeuralFDR and AdaFDR, to adaptively learn the optimal p-value threshold from covariates to significantly improve detection power while controlling the false discovery proportion.